Dr Steven Hussey

| Extra-ordinary Professor | |
Department |
|
Biochemistry, Genetics and Microbiology |
|
| This email address is being protected from spambots. You need JavaScript enabled to view it. |
Research Profiles
My Files
All of biology is transcriptionally wired. The regulation of secondary cell wall biosynthesis during wood formation in trees such as Eucalyptus is no exception. Previous studies, mostly in Arabidopsis and Populus, have deciphered a highly ramified transcriptional network hierarchically regulating secondary cell wall biosynthesis. Many of these transcription factors remain to be functionally characterized and their direct and indirect targets identified.
My group uses transcriptome data, DNA Affinity Purification sequencing (DAP-seq), transgenesis, protoplast transfection, bioinformatics and chromatin immunoprecipitation sequencing (ChIP-seq) to functionally characterize Eucalyptus transcription factors. Particularly, we are exploring recent high-throughput techniques for rapid and informative reconstruction of secondary cell wall transcriptional networks in non-model organisms. We are also interested in understanding the role of chromatin architecture in shaping transcriptional responses. In addition to understanding the fundamental biology of wood formation and how it is transcriptionally and epigenetically regulated, we hope to enhance economically important woody traits through manipulation of transcription factors regulating particular aspects of wood formation.

Transcriptional regulation and bioengineering of wood development group 2019. From left to right: Ms Ipeleng Randome (PhD student), Ms Ruby Ebbeling (BSc Hons student), Dr Raphael Ployet (postdoc), Dr Steven Hussey, Dr Letrisha Padayachee (postdoc), Ms Ingrid Roloff (BSc Hons student), Mr Lazarus Takawira (PhD student). In absentia: Ms Lungile Mabuza (MSc student), Ms Katrien Brown (MSc student).
Synthetic biology
The ability to re-engineer biological systems such as biochemical pathways or transcriptional networks is a hallmark of synthetic biology. We are starting to develop synthetic biology applications in forest trees, beginning with the design and chemical synthesis of 221 E. grandis secondary cell wall-related transcription factors and 65 promoters as standardized, domesticated Phytobricks that can be used for GoldenGate, MoClo and GoldenBraid 2.0 assembly. This work is part of a community DNA synthesis grant awarded by the DOE-JGI, and the synthetic panel will be freely available to the international community (see FMG's synthetic biology page).
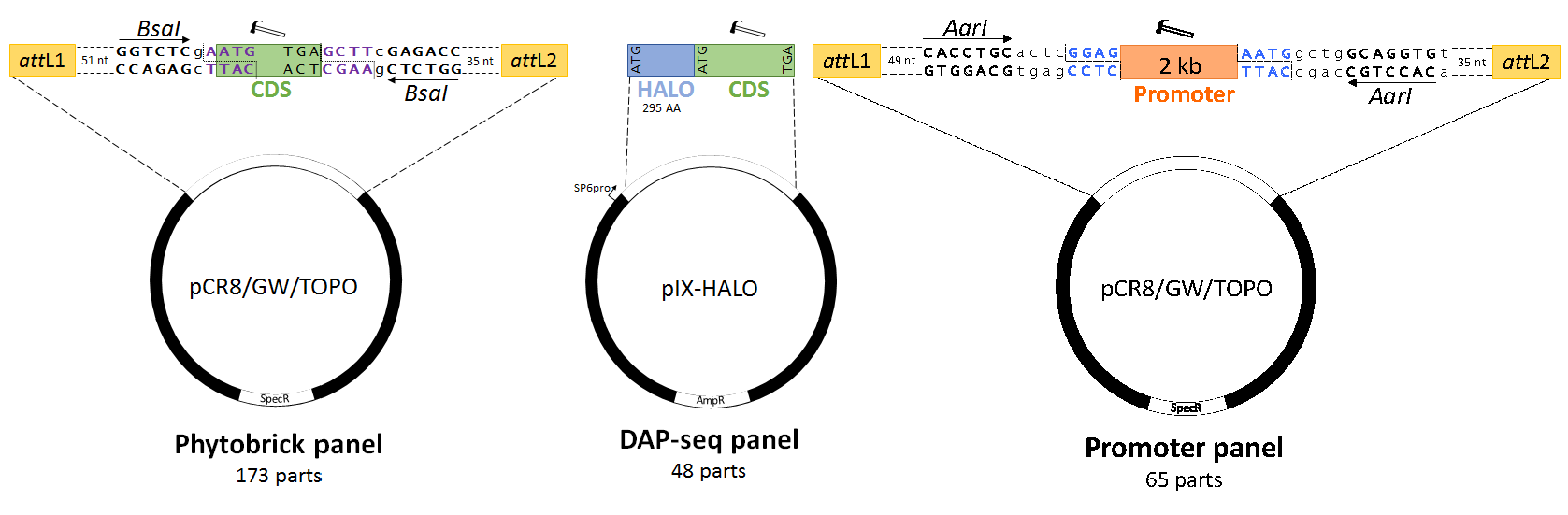
Figure 1. Design of standardized synthetic E. grandis SCW-related transcription factors and promoters (Hussey et al. 2019). Hammers indicate domesticated sequences, dotted lines indicate restriction cleavage sites, coloured bases indicate standard syntax sequences and lowercase sequences indicate spacer nucleotides. (A) Transcription factor Phytobricks contain attL GATEWAY recombination sites (green), fifty-one basepair chewback linkers, BsaI Type IIS recognition sites and standard syntax sequences (purple text). The start codon of the domesticated coding sequence (green) remains in frame with N-terminal tags in GATEWAY destination vectors, while being primarily intended for Golden Gate, MoClo and Golden Braid assembly. (B) The DNA Affinity Purification Sequencing (DAP-seq) panel of transcription factors is available as a C-terminal fusion to the HALO purification tag, intended for in vitro transcription and translation via the SP6 phage promoter. While not standardized by a universal syntax, the coding sequences are fully domesticated and can thus be subcloned as standardized parts into a universal acceptor plasmid. (C) The secondary cell wall promoter panel features semi-domesticated 2 kb promoter sequences (including 5’ UTRs) compatible with GATEWAY and AarI-mediated Golden Gate cloning. Standard prefix and suffix syntax sequences allow for two-step Type IIS assembly to any Phytobrick panel CDS in (A). AmpR, ampicillin resistance gene; CDS, coding sequence; SpecR, spectinomycin resistance gene.
Biomaker Africa
I am currently coordinating the Biomaker Programme at UP, in partnership with the UP Makerspace. UP has been awarded pump-priming support from the UK Global Challenges Research Fund through the University of Cambridge to build capacity in using Arduino-based plug-and-play hardware, low-cost open-source electronics and XOD visual programming for biological applications. Through the Biomaker model, multidisciplinary teams of varying levels of education and expertise are challenged to design and build practical devices, bioinstruments and biosensors using open-source parts and 3D printing. UP has held a number of training workshops and challenges in 2019, among them a two-day workshop run by Biomaker and plant synthetic biology leader Prof. Jim Haseloff. A number of Arduino-based training kits have been made available for student use at the UP Makerspace.
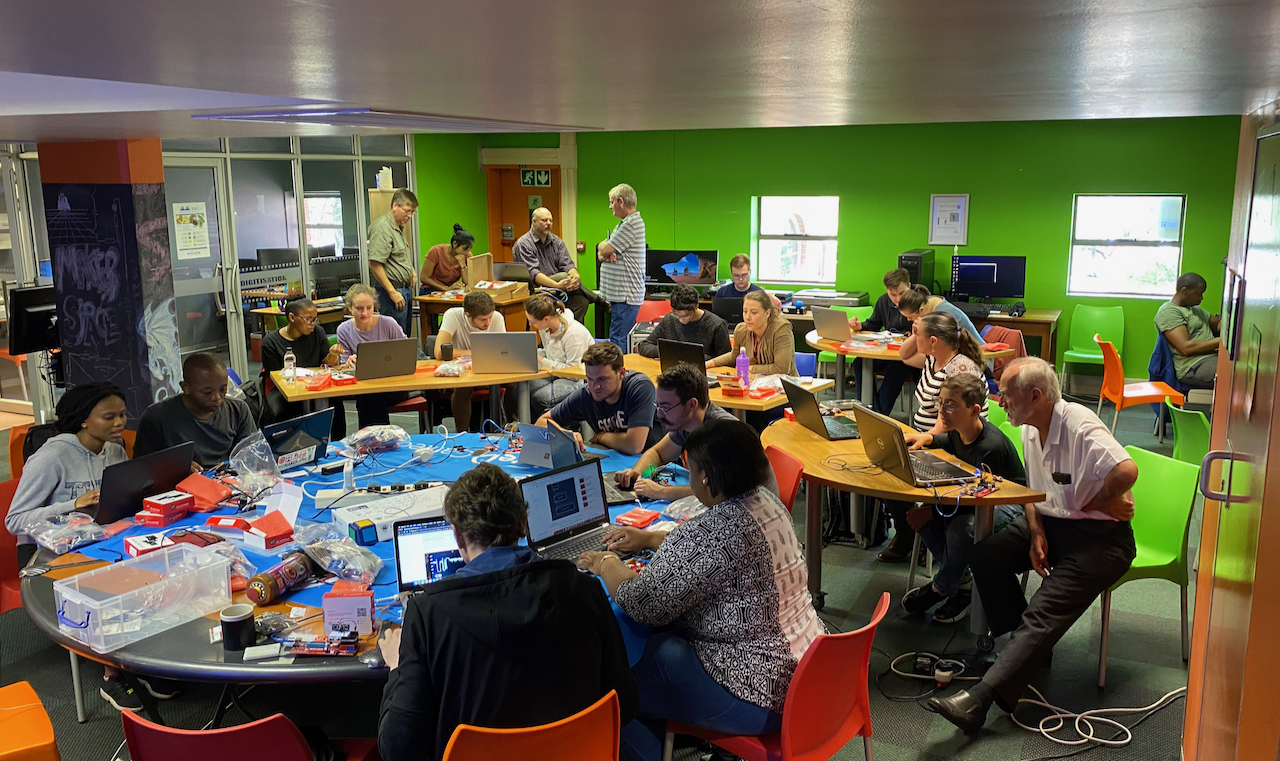


Low-cost viral diagnostics (LCVD) collaboration
I am also involved in research aimed developing new low-cost point-of-care diagnostics technology based on CRISPR/Cas, cell-free expression, isothermal nucleic acid amplification and fluorescent as well as lateral flow detection systems. These modular diagnostic assay components can be customized and optimized for wide disease detection applications in the biomedical, agricultural and forestry sciences. Dr Letrisha Padayachee, a postdoctoral fellow at UP working on efficient viral genomic DNA extraction and RPA-based target amplification, has also set up a protein expression platform based on bacterial cell-free extracts. The project, led by Prof. Jim Ajioka, is a collaborative effort with five PIs from the University of Cambridge, Imperial College London, the CSIR and UP. Together, they have run training workshops in viral diagnostics development at the Future Africa campus.

My Book Chapters
| Book Chapters |
|---|
| Hussey SG. 2022. Transcriptional regulation of secondary cell wall formation and lignification. In: Advances in Botanical Research. Academic Press, Cambridge, MA, pp . |
| Hussey SG, Wegrzyn JL, Vasquez-Gross HA. 2016. Evolutionary Histories of Gene Families in Angiosperm Trees. In: Comparative and Evolutionary Genomics of Angiosperm Trees. Groover A, Cronk Q. (eds). Springer International Publishing, Cham, pp 1-17. |
My Journal Articles
| Publication |
|---|
| Takawira LT, Mounet F, Mizrachi E, Myburg AA, Grima-Pettenati J, Rai A, Dupas A, Ladouce N, Christie N, San Clemente H, Thulloch J, Ployet R, Bachir H, Hussey SG. (2023) Functional investigation of five R2R3-MYB transcription factors associated with wood development in Eucalyptus using DAP-seq-ML. Plant Molecular Biology 113:33-57.
10.1007/s11103-023-01376-y |
| Karannagoda N, Spokevicius A, Hussey SG, Cassan-Wang H, Grima-Pettenati J, Bossinger B. (2022) Eucalyptus grandis AUX/INDOLE‑3‑ACETIC ACID 13 (EgrIAA13) is a novel transcriptional regulator of xylogenesis. Plant Molecular Biology
10.1007/s11103-022-01255-y |
| Karannagoda N, Spokevicius A, Hussey S, Bossinger G. (2020) Microanalytical techniques for phenotyping secondary xylem. IAWA Journal 41(3):356-389.
10.1163/22941932-bja10034 |
| Lyall R, Schlebusch SA, Proctor J, Prag M, Hussey SG, Ingle RA, Illing N. (2019) Vegetative desiccation tolerance in the resurrection plant Xerophyta humilis has not evolved through reactivation of the seed canonical LAFL regulatory network. The Plant Journal
10.1111/tpj.14596 |
| Tonfack LB, Hussey SG, Veale A, Myburg AA, Mizrachi E. (2019) Analysis of orthologous SECONDARY WALL-ASSOCIATED NAC DOMAIN1 (SND1) promotor activity in herbaceous and woody angiosperms. International Journal of Molecular Sciences 20:4623.
10.3390/ijms20184623 |
| Myburg AA, Hussey SG, Wang JP, Street NR, Mizrachi E. (2019) Systems and Synthetic Biology of Forest Trees: A Bioengineering Paradigm for Woody Biomass Feedstocks. Frontiers in Plant Biology 10:775.
10.3389/fpls.2019.00775 |
| Hussey SG, Grima-Pettenati J, Myburg AA, Mizrachi E, Brady SM, Yoshikuni Y, Deutsch S. (2019) A standardized synthetic Eucalyptus transcription factor and promoter panel for re-engineering secondary cell wall regulation in biomass and bioenergy crops. ACS Synthetic Biology 8(2):463-465.
10.1021/acssynbio.8b00440 |
| Brown K, Takawira L, O'Neill M, Mizrachi E, Myburg A, Hussey S. (2019) Identification and functional evaluation of accessible chromatin associated with wood formation in Eucalyptus grandis. New Phytologist 223(4):1937-1951.
10.1111/nph.15897 |
| Laubscher M, Brown K, Tonfack LB, Myburg AA, Mizrachi E, Hussey SG. (2018) Temporal analysis of Arabidopsis genes activated by Eucalyptus grandis NAC transcription factors associated with xylem fibre and vessel development. Scientific Reports 8:10983.
10.1038/s41598-018-29278-w |
| Lu Q, Shao F, Macmillan C, Wilson IW, van der Merwe K, Hussey SG, Myburg AA, Dong X, Qiu D. (2018) Genomewide analysis of the lateral organ boundaries domain (LBD) gene family in Eucalyptus grandis reveals members that differentially impact secondary growth. Plant Biotechnology Journal
10.1111/pbi.12754 |
| Hussey SG, Loots MT, van der Merwe K, Mizrachi E, Myburg AA. (2017) Integrated analysis and transcript abundance modelling of H3K4me3 and H3K27me3 in developing secondary xylem. Scientific Reports 7:3370.
10.1038/s41598-017-03665-1 |
| Hussey SG, Mizrachi E, Groover A, Berger DK, Myburg AA. (2015) Genome-wide mapping of histone H3 lysine 4 trimethylation in Eucalyptus grandis developing xylem. BMC Plant Biology 15:117.
10.1186/s12870-015-0499-0 |
| Hussey SG, Saïdi MN, Hefer C, Myburg AA, Grima-Pettenati J. (2014) Structural, evolutionary and functional analysis of the NAC domain protein family in Eucalyptus. New Phytologist 206(4):1337-1350.
10.1111/nph.13139 |
| Myburg AA, Grattapaglia D, Tuskan GA, Hellsten U, Hayes RD, Grimwood J, Jenkins J, Lindquist E, Tice H, Bauer D, Goodstein DM, Dubchak I, Poliakov A, Mizrachi E, Kullan ARK, Hussey SG, Pinard D, Van der Merwe K, Singh P, et al. (2014) The genome of Eucalyptus grandis. Nature 510:356-362.
10.1038/nature13308 |
| Hussey SG, Mizrachi E, Creux NM, Myburg AA. (2013) Navigating the transcriptional roadmap regulating plant secondary cell wall deposition. Frontiers in Plant Science 4(325):1-21.
10.3389/fpls.2013.00325 |
| Hussey SG, Mizrachi E, Spokevicius AV, Bossinger G, Berger DK. (2011) SND2, a NAC transcription factor gene, regulates genes involved in secondary cell wall development in Arabidopsis fibres and increases fibre cell area in Eucalyptus. BMC Plant Biology 11:173.
10.1186/1471-2229-11-173 |
| Botha J, Pinard D, Creux N, Hussey S, Maritz-Olivier C, Spokevicius A, Bossinger G, Mizrachi E, Myburg AA. (2011) Characterising the role of the Eucalyptus grandis SND2 promoter in secondary cell wall biosynthesis. BMC Proceedings 5(7):P105.
10.1186/1753-6561-5-S7-P105 |
| Hussey S, Mizrachi E, Berger D, Myburg AA. (2011) The role of SND2 in the regulation of Arabidopsis fibre secondary cell wall formation. BMC Proceedings 5(7):P114.
10.1186/1753-6561-5-S7-P114 |

